|
This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
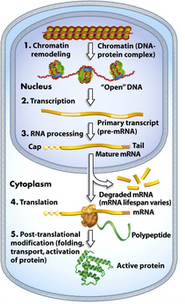
What is gene expression? Every cell in our body contains the same genetic material, or DNA: skin cells contain the same DNA as liver cells, cornea cells, and heart cells just to give an example. So if it is not differences in DNA content in each cell that makes them what they are there must be something else at play. In fact, it is the expression of DNA that causes skin cells to be skin and liver cells to be liver. Genes can be turned on and off through regulatory mechanisms illustrated on the right. These mechanisms include methylation pattens and histone modification, RNA processing, and post translational modification which allow gene products to be expressed or silenced. Genes are expressed in a cell specific manor and expression level can be influenced by many factors including disease. |
How is gene expression measured?
|
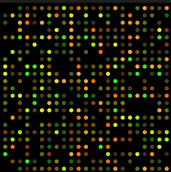
Gene expression can be measured using microarrays. Microarray technology, pictured left, measuring gene expression by using high-throughput screening via a small chip covered with microscopic DNA spots, known as probes (2). Each probe can bind to a specific sequence of DNA that it is complementary to in order to measure gene expression. The DNA segments of interest are labeled which chemicals such as fluorophores, indicated by the colors of the left, which allow for quantitative information on gene expression when microarrays are interpreted or read (2).
More information along with a microarray tutorial can be found here |
Gene Expression and COL3A1
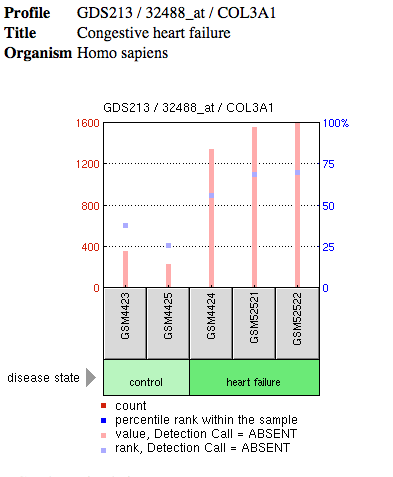
Searching COL3A1 on the GEO database yielded 8415 results. These resulting microarray experiments quantify expression levels of COL3A1 either after drug treatment or in a disease state. Many of the results were not relevant to EDS specifically and interestingly many involved looking at COL3A1 levels in individuals affected with HIV or cancer syndromes. One study that may be insightful towards our study of EDS quantified COL3A1 expression level in individuals with congenital heart failure.
|
Analysis: These findings which suggest that COL3A1 expression levels are higher in those with heart failure is interesting in the context of EDS. Males have been shown to have a higher level of collagen than females during adolescence. In addition, males with Vascular EDS die more than 2X more often than females during this time. Many of these deaths can be attributed to arterial rupture. It is also known that congestive heart failure can be due to arterial defects including plaque buildups and arterial blockage (1). In order to further explore the connection between collagen levels in healthy and diseased individuals, the same study could be done using healthy (control) and Vascular EDS affected individuals in both male and female populations. It would be interesting to determine of adolescent males with Vascular EDS express levels of COL3A similar to that of healthy males. If so, perhaps collagen levels as a cause of arterial disfunction and arterial rupture in the case of Vascular EDS can be further explored as a mortality mechanism and a relationship between COL3A1 levels and vascular EDS can be determined. |
Figure 1: Microarray data of COL3A1 expression levels in healthy individuals
compared to individuals with heart failure
compared to individuals with heart failure
References
1) "Heart Disease and Congestive Heart Failure." WebMD. Web. <http://www.webmd.com/heart-disease/guide-heart-failure#2>.
2) "DNA Microarray." Genetic Science Learning Center. University of Utah Health Sciences, n.d. Web. 25 Mar. 2015. <http://learn.genetics.utah.edu/content/labs/microarray/>.
1) "Heart Disease and Congestive Heart Failure." WebMD. Web. <http://www.webmd.com/heart-disease/guide-heart-failure#2>.
2) "DNA Microarray." Genetic Science Learning Center. University of Utah Health Sciences, n.d. Web. 25 Mar. 2015. <http://learn.genetics.utah.edu/content/labs/microarray/>.