This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
What is homology?
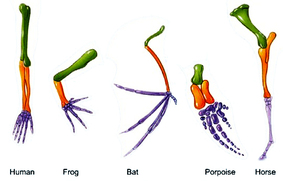
Homology is defined as shared ancestry of structures and genes that are seen between species (1). Homology arrises from common ancestors shared between species. Homologies can be quantified on both a molecular and phenotypic level. Pictured left are phenotypic homologies between the forearms of several different species. The similarity between the forearms of a human and a bat may seem surprising until we look deeper into the molecular basis of these similarities. By using an online tool called BLAST we can compare the DNA, mRNA, or proteins between different species. BLAST quantifies homology between species with percent identity which reflects the percent of the sequences that match between two species (2). Here protein sequences between different species were compared. Proteins are made up of different amino acid, and each amino acid has a 3 nucleotide code. When looking at homology of proteins between species you are looking at how well conserved the amino acid sequence between species are.
In order to gain a greater understanding of how conserved COL3A1 is among different species, reciprocal BLASTS were ran between the humans against the species below and percent identities were obtained.
Homo Sapiens (Human) COL3A1
Accession: NP_000081.1
Accession: NP_000081.1
|
Mus Musculus (Mouse) Col3a1
Accession: NP_034060.2 Percent Identity: 90% E-Value: 0.0 Rattus Norvegicus (Rat) Col3a1 Accession: NP_114474.1 Percent Identity: 90% E-Value: 0.0 Macaca Mulatta (Rhesus Macaque) COL3A1 Accession: NP_001252968.1 Percent Identity: 98% Evalue: 0.0 |
Pan Troglodytes (Chimpanzee) COL3A1
Accession: XP_001163809.1 Percent Identity: 100% E-Value: 0.0 Gallus Gallus (Chicken) COL3A1 Accession: NP_990711.2 Percent Identity: 72% E-Value: 0.0 |
Analysis
As stated on the gene homology page, the homology for proteins and DNA are different than one another because some of the nucleotides in DNA do not code for protein. Mice, the model organism in these studies actually has better protein homology with humans, 90%, compared to DNA homology of 86%. This illustrates the importance of looking into both protein and DNA homology when choosing a model organism.
References:
1) "Homologies." Understanding Eolution. University of California Museum of Paleontology, Web. 25 Mar. 2015. <http://evolution.berkeley.edu/evolibrary/article/lines_04>
2) Wheeler, David. BLAST QuickStart. U.S. National Library of Medicine, n.d. Web. 25 Mar. 2015. <http://www.ncbi.nlm.nih.gov/books/NBK1734/.>
3)"Basic Local Alignment Search Tool." BLAST Web. 23 Mar. 2015. <http://blast.ncbi.nlm.nih.gov/Blast.cgi>.
1) "Homologies." Understanding Eolution. University of California Museum of Paleontology, Web. 25 Mar. 2015. <http://evolution.berkeley.edu/evolibrary/article/lines_04>
2) Wheeler, David. BLAST QuickStart. U.S. National Library of Medicine, n.d. Web. 25 Mar. 2015. <http://www.ncbi.nlm.nih.gov/books/NBK1734/.>
3)"Basic Local Alignment Search Tool." BLAST Web. 23 Mar. 2015. <http://blast.ncbi.nlm.nih.gov/Blast.cgi>.
