This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
What is Proteomics?
Proteomics is the large scale study or proteomes, or total collection of proteins in an organism (7). Information obtained from proteomics includes function, structure, interactions, and modifications. Here we will take a closer look at COL3A1's domains and motifs, interaction partners, and post translational modifications to gain a better understanding of how mutations in COL3A1 lead to Vascular EDS.
What are Domains and Motifs?
|
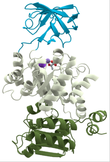
Domains are discrete functional units of proteins that are often conserved (1). Domains, which fold independently and have their own function, range in length from 25-500 amino acids long can be looked at as molecular building blocks for overall protein function. For example, three distinct domains pictured left which make up Pyruvate Kinase are depicted in different colors to demonstrate their independent folding patterns and unique functions. Motifs are short highly conserved regions arising from secondary structures including alpha-helixes and beta sheets. Motifs that give rise to domains and are often critical to domain function such as active sites or nuclear localization sequences (1).
What are the domains of COL3A1? |
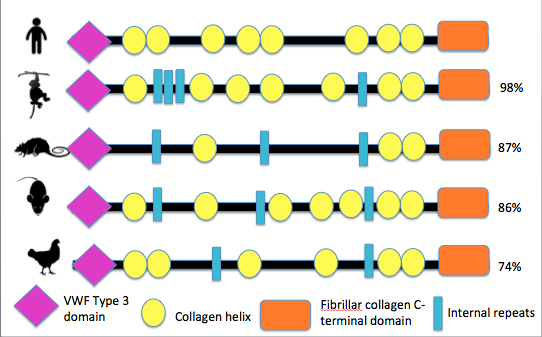
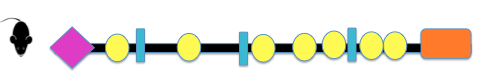
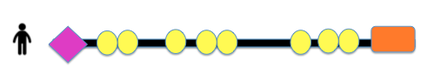
The domains of COL3A1 in various organisms, pictured below, were compiled using PHAM and SMART. The resulting domains include the Von Willebrand factor, Type C, Collagen helix, and Fibrillar collagen C-terimnal. Von Willebrand factor, Type C (VWF) is often involved in the formation of multicellular complexes via cell adhesion, migration, signal transduction, and other biological processes (2). Collagen helix domains, found in collagen molecules, are GLY-X-Y repeats that form a triple helix. 2/3 of known Vascular EDS mutation are Glycine substitutions in this domain. Fibrillar collagen C-terminal is found at the terminal of fibrillar collagens such as COL3A1.
Figure 1: Schematic representation of COL3A1 domain across different speices
Analysis
COL3A1 domains are highly conserved among different species indicating the importance of the VWF Type 3, Collagen helix, and Fibrillar collagen-C terminal domains to the function of the protein in many different organisms. Interestingly, the glycine substitution mutations that have been documented to account for 2/3 of Vascular EDS, are in a collagen helix domain not identified by PHAM or SMART. This highlights the importance of consulting multiple databases when gathering information regarding protein domains.
What are protein interaction networks?
Protein interaction networks are defined as complexes serving a distinct biological process that are connected via biochemical interactions and electrostatic forces (4). When understanding a disease and its mechanisms, it is very important to look beyond the directly mutated gene. Because of the interconnection of pathways and networks in our bodies, one mutation gene in single gene diseases such as Vascular EDS can have a cascade of effects, resulting in different function for many other proteins and pathways. By gaining an understanding of these interactions, new avenues for therapies and treatment can be explored.
What proteins does COL3A1 interact with?
Analysis
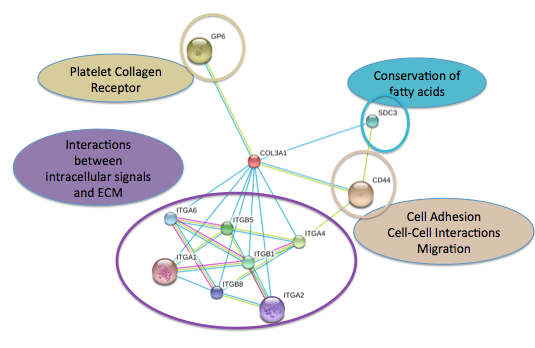
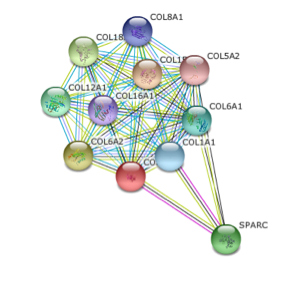
Figure 2 was obtained from SMART database and Figure 3 was obtained from String database. Figure 3 shows that COL3A1's closest interactions are to other collagens which is consistent with what we would expect as collagens often act together. SPARC, the other interaction partner in this network is responsible for cell growth and will prove to be important in future studies (see conclusions and future directions) (5).
Figure 2 shows the interaction partners beyond other collagens which reflects the importance of COL3A1 in many different cellular processes which I have highlighted in the figure. It is important to note that SPARC is not shown as an interaction partner in this network. This inconsistency can be potentially explained by what information each database has incorporated into their analysis. Even on a single database, individual networks are always changing as new research is assimilated. This highlight the importance of incorporating multiple databases into any analysis to obtain the most reliable and up to date information.
Figure 2 shows the interaction partners beyond other collagens which reflects the importance of COL3A1 in many different cellular processes which I have highlighted in the figure. It is important to note that SPARC is not shown as an interaction partner in this network. This inconsistency can be potentially explained by what information each database has incorporated into their analysis. Even on a single database, individual networks are always changing as new research is assimilated. This highlight the importance of incorporating multiple databases into any analysis to obtain the most reliable and up to date information.
What are post translational modifications?
|
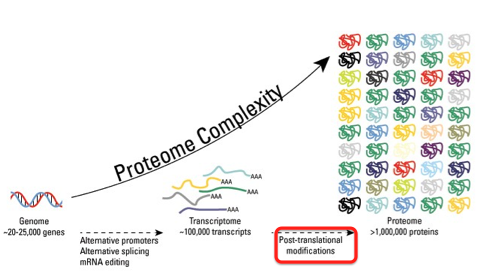
Post translational modifications are the final step in the central dogma turning proteins that have been translated into functional products. Post translational modifications are performed by enzymes and result in chemical modifications that can change the structure, function, or abundance of proteins. These chemical modifications include phosphorylation, glycoylation, ubiqitination, nitrosylation, methylation, acetylation, and lipidation (6) . Overall post translational modifications increase proteome complexity, as seen to the right. Post translational modifications can be measured using mass spectrometry, eastern blotting, or western blotting.
|
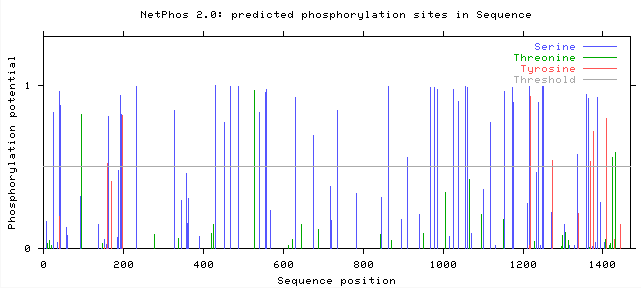
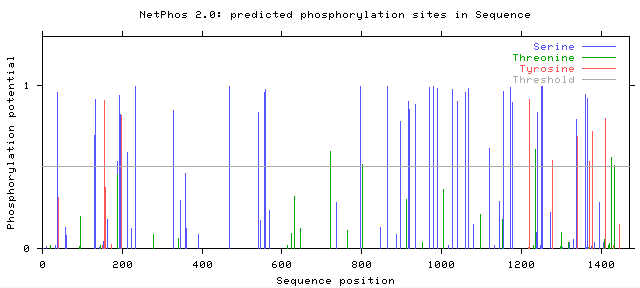
The post translational modification, phosphorylation, can be predicted using the bioinformatic tool Net Phos 2.0 which predicts the likelihood that Serine, Threonine, or Tyrosine shown in blue, green, and red respectively in the figures below will be phosphorylated. Figures 1 and 2 below show the predicted phosphorylation of human and mouse COL3A1. Lines that surpass the grey horizontal line through the middle, or "threshold", indicate that phosphorylation events at these sights have a greater than 50% chance of occurring.
Where does phosphorylation occur in mouse and human COL3A1?
Analysis
Both mouse and human COL3A1 have many sites with predicted Serine phosphorylation greater than 50%. Specifically mice have 37 sites and humans have 34 sites where Serine, Tyrosine, or Threonine have a greater than 50% chance of being phosphorylated . While the protein homology is only 90% between mice and humans, it is encouraging that their predicted phosphorylation patterns are similar because of the downstream effects that phosphorylation has on protein function. Thus, it is important to look beyond the amino acid sequence when comparing the similarity of genes and protein products in different organisms. In order to determine the importance of each phosphorylation site in the function of COL3A1, each site of phosphorylation could be replaced with an animo acid that has similar properties but cannot be phosphorylated. This experiment could be done in mice and would provide more insights on the importance of phosphorylation of each amino acid in COL3A1 function and disease mechanism.
References:
1) Stanley, Pamala. "Protein Domains and Motifs." Protein Domains and Motifs - Pamela Stanley Lab Wiki. N.p., Mar. 2005. Web. 25 Mar. 2015. <http://stanxterm.aecom.yu.edu/wiki/index.php?page=Protein_domains_and_motifs>.
2) Colombatti A, Bonaldo P, Doliana R (1993) "Type A modules: interaction domains found in several non-fibrillar collagens and in other extacellular matrix proteins." Matrix 13 (4): 297-306
3) "Pfam: Home Page." Pfam: Home Page. Web. 25 Mar. 2015. <http://pfam.sanger.ac.uk/>
4) "Protein-Protein Interaction Networks." Nature.com Subject Areas. Nature Publishing Group, Web. 12 May 2015. <http%3A%2F%2Fwww.nature.com%2Fsubjects%2Fprotein-protein-interaction-networks>.
5) "SPARC Gene." - GeneCards. Web. 12 May 2015. <http://www.genecards.org/cgi-bin/carddisp.pl?gene=SPARC>.
6)"Overview of Post-Translational Modifications (PTMs)." Life Technologies. Thermo Fisher Scientific. Web. 12 May 2015. <https://www.lifetechnologies.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html>.
7) "Proteomics." American Medical Association, Web. 12 May 2015. <http://www.ama-assn.org/ama/pub/physician-resources/medical-science/genetics-molecular-medicine/current-topics/proteomics.page>.
1) Stanley, Pamala. "Protein Domains and Motifs." Protein Domains and Motifs - Pamela Stanley Lab Wiki. N.p., Mar. 2005. Web. 25 Mar. 2015. <http://stanxterm.aecom.yu.edu/wiki/index.php?page=Protein_domains_and_motifs>.
2) Colombatti A, Bonaldo P, Doliana R (1993) "Type A modules: interaction domains found in several non-fibrillar collagens and in other extacellular matrix proteins." Matrix 13 (4): 297-306
3) "Pfam: Home Page." Pfam: Home Page. Web. 25 Mar. 2015. <http://pfam.sanger.ac.uk/>
4) "Protein-Protein Interaction Networks." Nature.com Subject Areas. Nature Publishing Group, Web. 12 May 2015. <http%3A%2F%2Fwww.nature.com%2Fsubjects%2Fprotein-protein-interaction-networks>.
5) "SPARC Gene." - GeneCards. Web. 12 May 2015. <http://www.genecards.org/cgi-bin/carddisp.pl?gene=SPARC>.
6)"Overview of Post-Translational Modifications (PTMs)." Life Technologies. Thermo Fisher Scientific. Web. 12 May 2015. <https://www.lifetechnologies.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html>.
7) "Proteomics." American Medical Association, Web. 12 May 2015. <http://www.ama-assn.org/ama/pub/physician-resources/medical-science/genetics-molecular-medicine/current-topics/proteomics.page>.